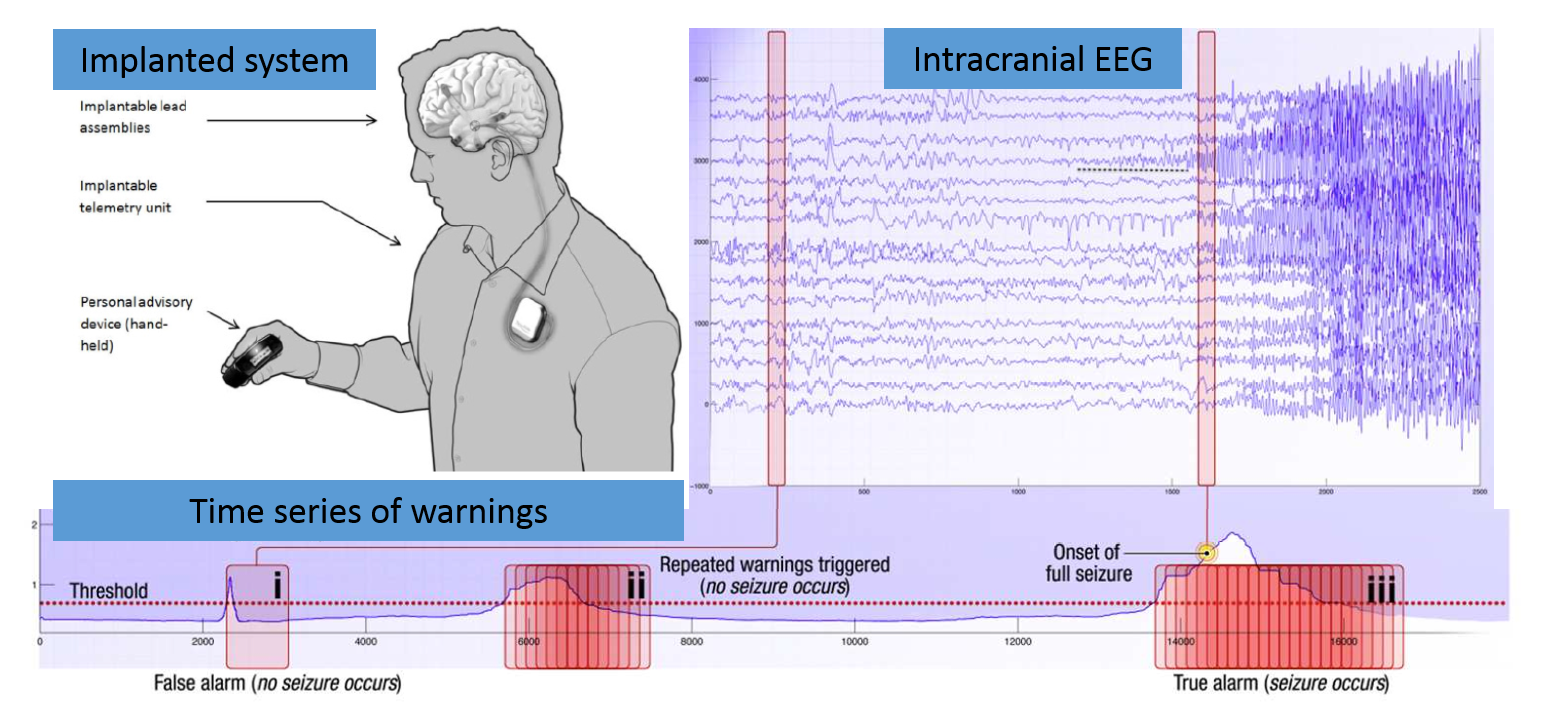
How it works
Epilepsyecosystem shares world-class datasets for epilepsy research with a special focus on facilitating international-scale interaction to develop, crowdsource and share seizure prediction algorithms. We currently offer access to two kinds of datasets. When you register with Epilepsyecosystem you gain access to these datasets under the terms and conditions specified. Click the links to find out out more:
My Seizure Gauge data for non-invasive wearables based seizure prediction.